This function calculates smoothed estimates of circular standard deviation using weighted kernel density estimation. It works similarly to density_asymmetry by applying Gaussian weights across an x-axis variable to compute localized circular SD estimates.
Usage
smoothed_circ_sd(
dt,
circ_space = 180,
weights_sd = 10,
xvar = "abs_td_dist",
yvar = "err",
by = c(),
n_points = 91
)Arguments
- dt
data.table with the data.
- circ_space
Circular space, which can be 180 or 360 (default: 180).
- weights_sd
Standard deviation of the Gaussian window to use across
xvar(default: 10).- xvar
X-axis variable, such as dissimilarity between items (default: "abs_td_dist").
- yvar
Y-axis variable, normally errors (default: "err").
- by
A vector of grouping variable names (default: an empty vector).
- n_points
The number of points along the x-axis at which to compute circular SD (default: 91).
Value
A data.table with the grouping variables, dist - the values of X-axis variable at which the circular SD is computed, and circ_sd - the smoothed circular standard deviation estimate at each point.
Examples
data(Pascucci_et_al_2019_data)
ex_data <- Pascucci_et_al_2019_data
ex_data[, err := angle_diff_180(reported, orientation)] # response errors
#> observer orientation reported rt err
#> <int> <int> <int> <num> <num>
#> 1: 1 135 137 1.0829786 2
#> 2: 1 65 56 0.9887931 -9
#> 3: 1 61 65 1.5067748 4
#> 4: 1 27 25 1.9070205 -2
#> 5: 1 22 20 2.0247443 -2
#> ---
#> 4396: 10 35 26 1.7775651 -9
#> 4397: 10 141 135 2.0365374 -6
#> 4398: 10 178 163 1.1301296 -15
#> 4399: 10 168 168 1.3772832 0
#> 4400: 10 24 28 2.3897599 4
ex_data[, prev_ori := shift(orientation), by = observer] # orientation on previous trial
#> observer orientation reported rt err prev_ori
#> <int> <int> <int> <num> <num> <int>
#> 1: 1 135 137 1.0829786 2 NA
#> 2: 1 65 56 0.9887931 -9 135
#> 3: 1 61 65 1.5067748 4 65
#> 4: 1 27 25 1.9070205 -2 61
#> 5: 1 22 20 2.0247443 -2 27
#> ---
#> 4396: 10 35 26 1.7775651 -9 68
#> 4397: 10 141 135 2.0365374 -6 35
#> 4398: 10 178 163 1.1301296 -15 141
#> 4399: 10 168 168 1.3772832 0 178
#> 4400: 10 24 28 2.3897599 4 168
# determine the shift in orientations between trials
ex_data[, diff_in_ori := angle_diff_180(prev_ori, orientation)]
#> observer orientation reported rt err prev_ori diff_in_ori
#> <int> <int> <int> <num> <num> <int> <num>
#> 1: 1 135 137 1.0829786 2 NA NA
#> 2: 1 65 56 0.9887931 -9 135 70
#> 3: 1 61 65 1.5067748 4 65 4
#> 4: 1 27 25 1.9070205 -2 61 34
#> 5: 1 22 20 2.0247443 -2 27 5
#> ---
#> 4396: 10 35 26 1.7775651 -9 68 33
#> 4397: 10 141 135 2.0365374 -6 35 74
#> 4398: 10 178 163 1.1301296 -15 141 -37
#> 4399: 10 168 168 1.3772832 0 178 10
#> 4400: 10 24 28 2.3897599 4 168 -36
ex_data[, abs_diff_in_ori := abs(diff_in_ori)]
#> observer orientation reported rt err prev_ori diff_in_ori
#> <int> <int> <int> <num> <num> <int> <num>
#> 1: 1 135 137 1.0829786 2 NA NA
#> 2: 1 65 56 0.9887931 -9 135 70
#> 3: 1 61 65 1.5067748 4 65 4
#> 4: 1 27 25 1.9070205 -2 61 34
#> 5: 1 22 20 2.0247443 -2 27 5
#> ---
#> 4396: 10 35 26 1.7775651 -9 68 33
#> 4397: 10 141 135 2.0365374 -6 35 74
#> 4398: 10 178 163 1.1301296 -15 141 -37
#> 4399: 10 168 168 1.3772832 0 178 10
#> 4400: 10 24 28 2.3897599 4 168 -36
#> abs_diff_in_ori
#> <num>
#> 1: NA
#> 2: 70
#> 3: 4
#> 4: 34
#> 5: 5
#> ---
#> 4396: 33
#> 4397: 74
#> 4398: 37
#> 4399: 10
#> 4400: 36
circ_sd_smooth <- smoothed_circ_sd(ex_data[!is.na(diff_in_ori)],
circ_space = 180, weights_sd = 10, xvar = "abs_diff_in_ori",
yvar = "err", by = c("observer")
)
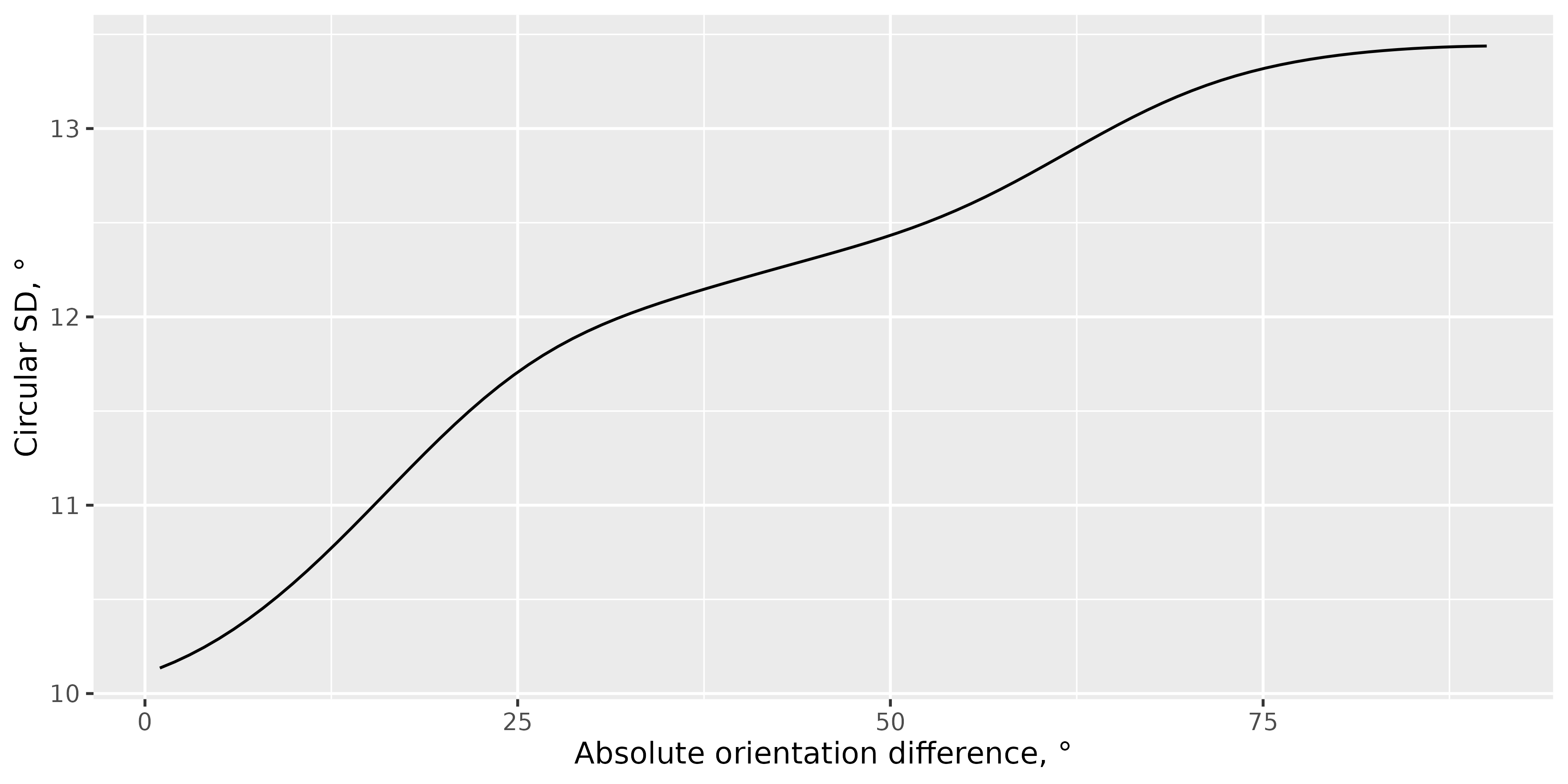
library(ggplot2)
ggplot(circ_sd_smooth, aes(x = dist, y = circ_sd)) +
geom_line(stat = "summary", fun = mean) +
labs(y = "Circular SD, °", x = "Absolute orientation difference, °")